Introduction
The data made available here was collected and analysed in a large genome-wide association study performed with more than 2000 Crl:CFW(SW)-US_P08 outbred mice and presented in the 2016 articles:
- “Genome-wide association of multiple complex traits in outbred mice by ultra low-coverage sequencing” by Nicod et al. (Free read-only link. View Abstract
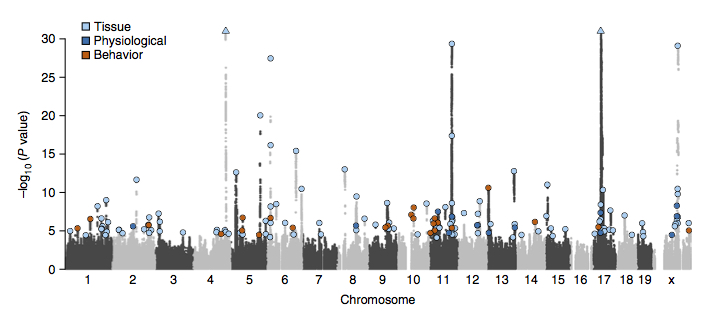
Two bottlenecks impeding the genetic analysis of complex traits in rodents are access to mapping populations able to delivergene-level mapping resolution and the need for population-specific genotyping arrays and haplotype reference panels. Here we combine low-coverage (0.15×) sequencing with a new method to impute the ancestral haplotype space in 1,887 commercially available outbred mice. We mapped 56 unique quantitative trait loci for 92 phenotypes at a 5% false discovery rate. Gene-level mapping resolution was achieved at about one-fifth of the loci, implicating Unc13c and Pgc1a at loci for the quality of sleep, Adarb2 for home cage activity, Rtkn2 for intensity of reaction to startle, Bmp2 for wound healing, Il15 and Id2 for several T cell measures and Prkca for bone mineral content. These findings have implications for diverse areas of mammalian biology and demonstrate how genome-wide association studies can be extended via low-coverage sequencing to species with highly recombinant outbred populations.
- “Rapid genotype imputation from sequence without reference panels” by Davies et al ( Free read-only link.View Abstract
Inexpensive genotyping methods are essential for genetic studies requiring large sample sizes. In human studies, array-based microarrays and high-density haplotype reference panels allow efficient genotype imputation for this purpose. However, these resources are typically unavailable in non-human settings. Here we describe a method (STITCH) for imputation based only on sequencing read data, without requiring additional reference panels or array data. We demonstrate its applicability even in settings of extremely low sequencing coverage, by accurately imputing 5.7 million SNPs at a mean r-squared value of 0.98 in 2,073 outbred laboratory mice (0.15× sequencing coverage). In a sample of 10,670 Han Chinese (1.7× coverage), we achieve accuracy similar to that of alternative approaches that require a reference panel, demonstrating that our approach can work for genetically diverse populations. Our method enables straightforward progression from low-coverage sequence to imputed genotypes, overcoming barriers that at present restrict the application of genome-wide association study technology outside humans.
- Additional data for the Micronucleus phenotype was generated for a separate publication “A genome-wide association study for regulators of micronucleus formation in mice” by McIntyre et al. and is also included here. View Abstract
In mammals the regulation of genomic instability plays a key role in tumor suppression and also controls genome plasticity, which is important for recombination during the processes of immunity and meiosis. Most studies to identify regulators of genomic instability have been performed in cells in culture or in systems that report on gross rearrangements of the genome, yet subtle differences in the level of genomic instability can contribute to whole organism phenotypes such as tumour predisposition. Here we performed a genome-wide association study in a population of 1379 outbred Crl:CFW(SW)-US_P08 mice to dissect the genetic landscape of micronucleus formation, a biomarker of chromosomal breaks, whole chromosome loss, and extra-nuclear DNA. Variation in micronucleus levels is a complex trait with a genome-wide heritability of 53.1%. We identify seven loci influencing micronucleus formation (false discovery rate<5%), and define candidate genes at each locus. Intriguingly at several loci we find evidence for sex-specific genetic effects in micronucleus formation, with a locus on chromosome 11 being specific to males.
- See also the paper “Genome-wide association study of behavioral, physiological and gene expression traits in outbred CFW mice” by Parker et al.
Phenotypes
Data available in the phenotypes sub-directory.
- TableS1_phenotypes.xlsx, also appearing in the article, describes the 200 phenotypes collected in this study. We report the number of mice generating data for the analysis (after exclusion of outliers), the mean and standard deviation for all animals and for males and females separately, the linear model used to generate residuals, the estimated heritability (with standard error and p-value) and to which category the phenotype belongs. The covariates used in the linear models are described at the bottom of the main table.
- CFW_measures.txt contains all 200 measures collected from the CFW mice.
- CFW_residuals.txt file contains the residuals calculated from the lineal model described in TableS1_phenotypes.xlsx
- CFW_covariates.txt contains all covariates collected during the study, including Sex. Their effect on each measure was assessed and if significant at P<-0.05 was included in a linear model to calculate residuals. If a covariate is used in a linear modes it is described at the bottom of the main table in TableS1_phenotypes.xlsx.
Genotype Dosages
Data available in the dosages and all_dosages sub-directories.
- dosages contain imputed allele dosages at the 359,559 tagging SNPs used for mapping. There is one .RData file per chromosome (e.g. chr19.prunedgen.final.maf001.0.98.RData). Each file contains 3 R objects:
- nameList: List with the IDs of the 2073 mice with imputed dosages. Note the format of the IDs (e.g. “Q_CFW-SW___100.0a_recal.bam”) differs from that used in the phenotypes files (“Q_CFW-SW/100.0a”).
- pruned_pos: Table with the SNP position and the reference (“REF”) and alternative (“ALT”) alleles.
- pruned_dosages: Table with the imputed allele dosages at the tagging SNPs for the 2073 mice.
- These files include dosages for all 2073 mice. In this directory is the file “List_of_1934_mice_used_for_analysis.RData” which contains the list of the 1934 mice retained after exclusion of related animals and outliers, and used for the genetic mapping.
- all_dosages contains imputed allele dosages at 7M sites. Data for each chromosome is stored as an Rdata file (eg chr19.dosages.RData)
- Each file contains a dataframe named df.
- Each row corresponds to one SNP.
- The first five columns of the dataframe are chr, bp, hwe, info, pass that give respectively the chromosome, base-pair position (build 38), Hardy-Weinberg Equilibrium p-value, info-score and QC pass/fail indicator (those that pass correspond to the 5.7M SNPs used for fine mapping, and are defined to be info>0.4 & hwe>1e-6 on autosomes and info>0.4 on chrX).
- The remaining columns give the imputed dosages for the 2073 mice (one column per mouse).
- Thus each dataframe has over 2000 columns and tens or hundreds of thousands of rows and is therefore VERY LARGE.
QTL Mapping results
Data available in the mapping sub-directory.
Association results (as -log10( P-values) ) are provided at 359,559 tagging SNPs for 200 phenotypes (one .txt file per phenotype). These data can be visualised at GScanViewer.
The sub directories mapping/micronucleus_males_only/ and mapping/micronucleus_females_only/ contain the mapping results of the Micronucleus.Mn.NCE phenotype performed separately with males and females.
QTL plots
Plots of all QTLs are available as .pdf files in the qtls sub-directory. Example plot
BAM files
The BAM files containing low-coverage whole genome sequence data for the CFW mice are available from the European Nucleotide Archive under project ERP001040 .
STITCH
The STITCH algorithm used to impute genotypes in this study was written by Robert Davies and is available from here.
Credits
The data was generated by or with the assistance of: Jerome Nicod, Robert Davies, Na Cai, Carl Hassett, Leo Goodstadt, Cormac Cosgrove, Benjamin K Yee, Vikte Lionikaite, Rebecca E McIntyre, Carol Ann Remme, Elisabeth M. Lodder, Jennifer S. Gregory, Tertius Hough, Russell Joynson, Hayley Phelps, Barbara Nell, Clare Rowe, Joe Wood, Alison Walling, Nasrin Bopp, Amarjit Bhomra, Polinka Hernandez-Pliego, Jacques Callebert, Richard M. Aspden, Nick P Talbot, Peter A Robbins, Mark Harrison, Martin Fray, Jean-Marie Launay, Yigal M. Pinto, David A. Blizard, Connie R. Bezzina, David J Adams, Paul Franken, Tom Weaver, Sara Wells, Steve DM Brown, Paul K Potter, Paul Klenerman, Arimantas Lionikas, Richard Mott & Jonathan Flint
Any queries should be addressed to Jerome Nicod, Richard Mott or Jonathan Flint .